Russian translation courtesy of Kate Osipova of Science Translations.

I started working on this project in the context of the larger Parametric Human Project during my time as a research intern at Autodesk Canada.
This project is very ambitious. The idea is to create a parametric model of the entire human skeleton to the greatest anatomical detail possible. We will collect a large number of digital samples and label the main anatomical landmarks. Using this information, along with some geometric analysis, we will align all the samples and place them into semantic correspondence. From this, we will build a statistical model of the geometry and apply it to a number of problems, such as bone identification and model-based image segmentation.
We are still in the initial stage of the project, determining the best approach to collecting a large number of digital bone samples. We have explored using medical image data, and computed tomography (CT) scans in particular, as our main source of data. CT scans are particularly useful for obtaining bone data since they highlight this particular tissue very well, and the exact (Houndsfield) values of the image pixel corresponding to bone are well known.
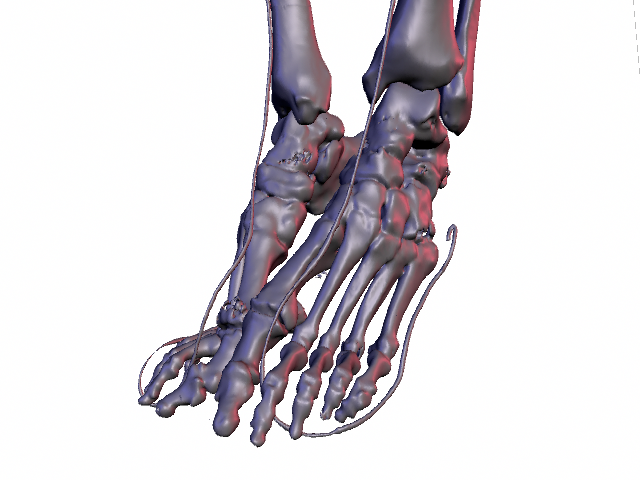
To extract the bone surfaces from CT, a standard thresholding or marching cubes algorithm is used. This produces results much like the image on the top of this page, which is showing the lower legs of the female So far, we have been able to solve the first problem by implementing the Interactive Watershed Transform (IWT). This algorithm is similar to the standard watershed transform but stores a record of new basins and basin merging points so that the final segmentation can be computed interactively from user placed markers. Our implementation runs on MeVisLab. Some of our results are shown below.
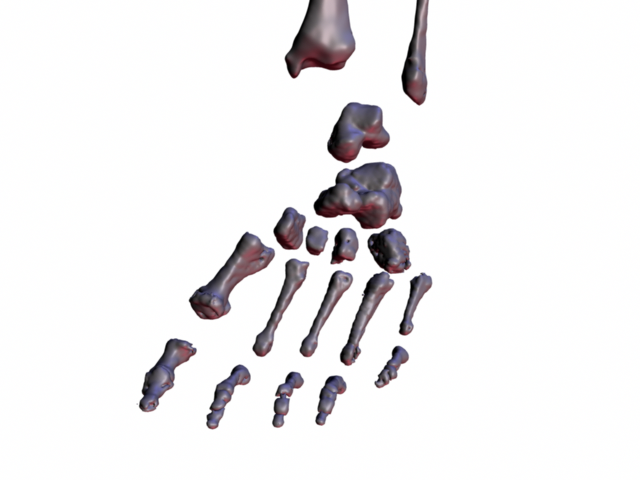
Individual Visible Female lower leg bones.
The main issue we have with this algorithm is that it is still dependent on the standard thresholding algorithm to produce the final segmentation result. Therefore, holes appear in areas where the bones are thin. This is a big issue near in the skull and certain regions at the ends of long bones.
Our current efforts are working toward fixing these issues. We are considering using structure tensor approaches to determine where the bones interface with soft tissue. We are also considering alternate sources of data, such as 3-D scans of cadaveric specimens.